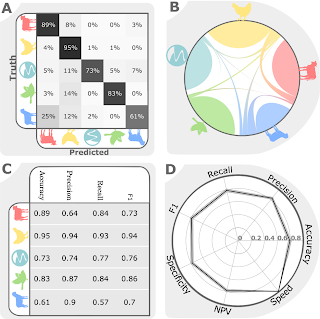
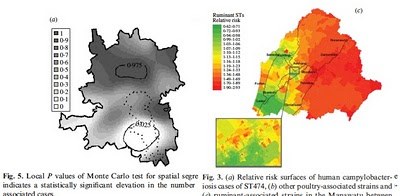
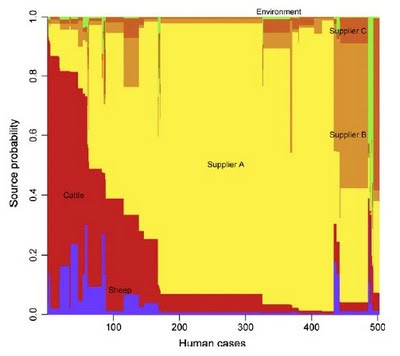
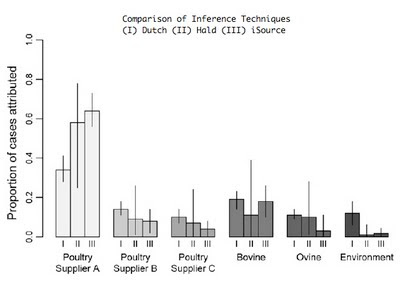
This study, published in October in PLOS Genetics, brings together machine learning, large bacterial isolate collections and whole genome sequencing to address the general problem of how to trace the source of human infections.
Specifically, we investigated campylobacteriosis, a common infection of animal origin causing ~1.5 million cases of gastroenteritis and 10,000 hospitalizations every year in the United States alone. We show that our combined machine learning/genomics analyses:
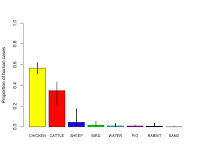
- Improve the accuracy with which infections can be traced back to farm reservoirs.
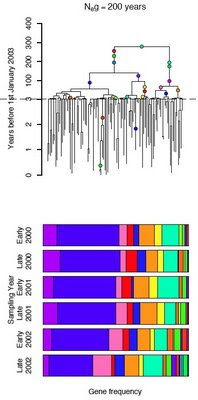
- Identify evolutionary shifts in bacterial affinity for livestock host species.
- Detect changes in human infection capability within related strains.
These results will improve understanding not only of Campylobacter, but more generally as these technologies can readily be applied to other important bacterial pathogen species.
This paper builds on previous work published by the group, including our well cited Tracing the source of campylobacteriosis (Wilson et al 2008, PLOS Genetics 4:e1000203). The use of these methods for tracing infection has influenced public health policy and contributed to reducing disease burden.
This work demonstrates the potential for modern genomics and artificial intelligence approaches to address common and serious problems that affect our everyday lives. The awareness of the importance of infection to society has rarely been higher than in 2021, and while the current pandemic imposes an acute global problem, other infections continue to present long-term threats to health and productivity.